 Gehman Lab analytics studio
Gehman Lab analytics studio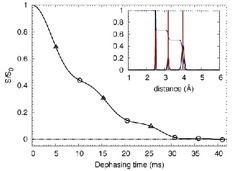
Boltzmann Statistics REDOR Analysis (BS-REDOR)
BS-REDOR employs Boltzmann maximum entropy statistics to reconstruct an unbiased distance distribution for
experimentally limited REDOR data sets. It was introduced in
Gehman et al. J Phys Chem B. 111(27):7802-11 (2007).
This page makes available for your REDOR analyses our current implementation of this theory.
An evolving guide to using the software can be downloaded here:
bsredor_manual.pdf, or as part of the current
distribution (version 1.0), available as a bzipped tarball here: bsredor01.tbz.
You probably also want to install the GNU Scientific Library and
gnuplot as discussed in the manual.
To compile this on your Windows machine we can suggest installing Cygwin - a Linux like environment for Windows.
We have successfully compiled the BS-REDOR code using cygwin, but remember to also install the GNU Scientific Library.
Boltzmann Statistics Magic Angle Spinning Sideband Analysis (BS-MAS)
As described in Sani et al.
Biophysical Journal 100:L40-L42 (2011). Software package is available here:
BS-MAS.tbz, and also from git:
BS-MAS.git
DePaking
Our implemention of McCabe and Wassall's FT de-Paking algorithm is provided here as an NMRPipe macro,
as described in Sani et al.
PeerJ 1:e30; DOI:10.7717/peerj.30 (2013). NMRPipe versions prior to 6.1 use the numeric parameter codes in
dePakeFT_pre61.M, while NMRPipe version 6.1 and later
includes dePakeFT.M by default. These scripts should be placed
with the other macros in $NMRTXT as dePakeFT.M (e.g. download, cd to download directory, and
using tcsh run `mv dePakeFT*.M $NMRTXT/dePakeFT.M).
You can also download a bzipped tar file (here: 2H_sw4M_d54DMPC_30C.tbz) which includes the same (Varian) 2H experimental data used in the paper, with the nmrPipe processing file. Please refer to the README.txt explanation file, also included.
enAMBlER
As in "enabler for AMBER" ... A few handy utilities we've developed to help with running AMBER. So far it's simply
a utilty written in Python to translate input file parameters into English. Find it at git:
enAMBlER.git.